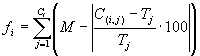One important application of GEP is symbolic regression or function finding, where the goal is to find a symbolic expression that performs well for all fitness cases within a certain error of the correct value. For most symbolic regression applications it is important to use small relative or absolute errors in order to discover a very good solution. But if we excessively narrow the range of selection and only allow the selection of individuals performing within a very small error, populations evolve very inefficiently and, most of the times, are incapable of finding a satisfactory solution. On the other hand, if we do the opposite and excessively enlarge the range of selection, numerous solutions with maximum fitness
fmax will appear that are far from good solutions. So, a simple fitness function which only discriminated between cases solved within and without the chosen error will not suffice, and a more skilled approach is required.
To solve this dilemma, I devised an evolutionary strategy that permits the discovery of very good solutions without halting evolution
(Ferreira 2001). With this scheme, the system is left to find for itself the best possible solution within a minimum error. For that purpose, a very broad limit for selection to operate is given, for instance, a relative error of 20% or 100%, that allows not only the evolutionary process to get started but also the fine tuning around the desired minimum error. And what is observed is that the individuals of early generations are usually very unfit but their later descendants are continually reshaped by the genetic operators, and populations adapt wonderfully, finding better solutions that progressively approach a very good solution. More formally, the fitness
fi of an individual program i is expressed by equation
(3.1a) if the error chosen is the absolute error, and by equation
(3.1b) if the error chosen is the relative error:
|
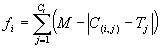
|
(3.1a) |
|
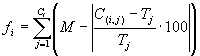
|
(3.1b) |
where M is the range of selection, C(i,j) the value returned by the individual program
i for fitness case j (out of Ct fitness cases) and
Tj is the target value for fitness case j. Note that the absolute value term corresponds, in both equations, to the error (absolute in the former, relative in the latter). This term is called precision
p. Thus, for a perfect fit, C(i,j)
= Tj and fi = fmax
= Ct*M.
We saw in section 3.1 another example of fitness function. In that case, the fitness of an individual was a function of the number of fitness cases correctly evaluated. For more complex Boolean applications, though, it is fundamental to penalize individuals able to solve correctly about 50% of fitness cases, as most probably this only reflects the 50% likelihood of correctly solving a binary Boolean function. So, it is advisable to select only individuals capable of solving more than 50-75% of fitness cases. Below that mark we can assign a symbolic value to fitness, for instance
fi = 1, or remove altogether these individuals from the populations by making them nonviable (fi = 0). Using this scheme, the process of evolution usually starts with very unfit individuals, as they are created very easily in initial populations. However, in future generations, highly fit individuals start to appear, rapidly spreading in the population. For such an easy problem as the Majority
(a, b, c) function analyzed in the previous section, this kind of scheme is not crucial, but for more complex problems it is convenient to choose a bottom line for selection. For those problems, the following fitness function can be used:
|
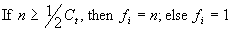
|
(3.2) |
where n is the number of fitness cases correctly evaluated, and
Ct is the total number of fitness cases.
Throughout this book we will come across other examples of fitness functions and we will see that the success of a problem greatly depends on the way the fitness function is designed. A classical example of a poorly designed fitness function is the one designed to predict the three-dimensional structure of proteins
(Schulze-Kremer 1992). Despite the high fitness of the evolved solutions, none of them was structurally similar to the target protein.
|