When a particular genome replicates itself and passes on the genetic information to the next generation, the sequence of the daughter molecule sometimes differs from that of the mother in one or more points. In spite of the virtual perfection of the replication machinery, sometimes a mismatched nucleotide is introduced in the newly synthesized strand. Although cells have mechanisms for correcting most mismatches (and for this, the complementary double-stranded DNA is extremely useful), some of them are not repaired and are passed on to the next generation
(Figure 1.3).
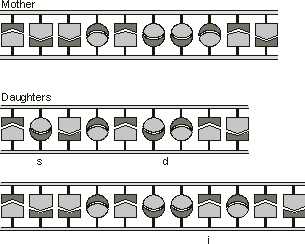
Figure 1.3. Mutations in the DNA sequence of genes. A base substitution
(s), a small deletion (d) and a small insertion (i) are shown here.
In nature, the rate at which mutation occurs is tightly controlled and different groups of organisms have different mutation rates, with virus and bacteria having higher mutation rates than eukaryotes. Of those, virus of course have the highest mutation rates as a single virion can leave hundreds or even thousands of progeny per infected cell, testing several new genomes in one generation.
It is important to analyze more closely the effects of point mutation on the protein itself. In a gene, the replacement of one nucleotide by another can have different effects: (1) the new codon might code for a new amino acid (these are called missense mutations); (2) the new codon might code for a “stop” codon, truncating the protein, or else a “stop” codon mutates into an amino acid codon, elongating the chain (nonsense mutations); (3) fairly frequently, though, point mutations in a gene have no effect at all in the protein sequence as the new codon might code for the same amino acid (neutral mutations); (4) in addition, in eukaryotes, where the sequence of most genes is interrupted by non-coding regions (introns), if a mutation occurs in an intron it has no effect whatsoever in protein sequence (also an example of a neutral mutation).
In another kind of protein mutation, large or small fragments may be inserted or deleted in the coding region of a gene. Large insertions or deletions almost invariably result in the production of a defective protein. The effect of short insertions or deletions depends on whether or not these modifications cause a shift in the reading frame of the gene (if they do, they are called frameshift mutations). If the fragment deleted or inserted is a multiple of three, then one or more codons are removed or added, resulting in the deletion or insertion of one or more amino acids in the protein. The consequences of these non-frameshift deletions/insertions are similar to the ones caused by missense mutations.
The effects of these kinds of mutations on the structure and functionality of a protein can be quite different. Point mutations may be neutral in effect, either not changing the amino acid at all or changing it by another that functions equally well in that position. The deletion/insertion of codons may also be of little consequence, changing only slightly the protein function. Occasionally, such mutations increase the efficiency of a protein, conferring some selective advantage for the organism itself. On the other hand, nonsense mutations and frameshift mutations have, almost every time, a lethal effect, especially if the new protein is fundamental to the survival of the organism. Nonetheless, very occasionally, such mutations might give rise to new, revolutionary traits.
We will see that, in GEP, most mutations, including point mutations and small insertions, have a profound effect in the structure and function of expression trees, more resembling the nonsense and frameshift mutations that occur in nature. Nonetheless, this type of mutation is extremely important in GEP evolvability and several new traits are introduced in this manner. However, less drastic mutations can also be found in GEP. As a matter of fact, some mutations change expression trees very smoothly and they might slightly or significantly increase the efficiency of the expression tree. Furthermore, in GEP, some mutations also have a clear neutral effect. For instance, mutations in the non-coding regions of genes have no effect whatsoever in the structure of expression trees. Other neutral mutations are not so easy to spot because they result in structurally different expression trees. In this case, the new expression tree is equivalent (in mathematical terms) to the parental expression tree. We will see that all kinds of mutation, from the most conservative to the most radical, are important to the evolution of good computer programs.
|
