We have seen that translation results in the formation of sub-ETs with different complexity, but the complete expression of the genetic information requires the interaction of these sub-ETs with one another. One of the simplest interactions is the linking of sub-ETs by a particular function. This process is similar to the assemblage of different protein subunits into a multi-subunit protein.
When the sub-ETs are algebraic or boolean expressions, any mathematical or boolean function with more than one argument can be used to link the sub-ETs into a final, multi-subunit ET. The functions most chosen are addition or multiplication for algebraic sub-ETs, and OR or IF for boolean
sub-ETs.
In the current version of GEP the linking function is a priori chosen for each problem, but it can be easily introduced in the
genome, for instance, in the last position of chromosomes, and also be subjected to adaptation. Indeed, preliminary results suggest that this system works very well.
Figure 3 illustrates the linking of two sub-ETs by addition.
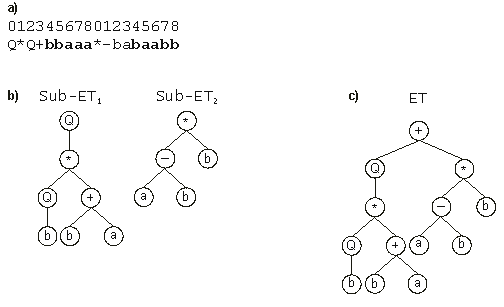
Figure 3. Expression of multigenic chromosomes as ETs. (a) A two-genic chromosome with the tails shown in bold.
(b) The sub-ETs codified by each gene. (c) The result of posttranslational linking with addition.
Note that the root of the final ET (+) is not encoded by the genome. Note also that the final ET could be linearly encoded as the following K-expression:
|
0123456789012 |
|
|
+Q**-bQ+abbba |
(3.10) |
However, to evolve solutions for complex problems, it is more effective
to use multigenic chromosomes, for they permit the modular construction of complex, hierarchical structures, where each gene codes for a small building block. These small building blocks are separated from each other, and thus can evolve independently. For instance, if we tried to evolve a solution for the symbolic regression problem presented in
section 6.1 with single-gene chromosomes, the success rate would fall significantly (see
section 6.1). In that case the discovery of small building blocks is more constrained as they are no longer free to evolve independently. This kind of experiment shows that GEP is in effect a powerful, hierarchical invention system capable of easily evolving simple blocks and using them to form more complex structures
[8, 9].
Figure 4 shows another example of sub-ET interaction, where three boolean sub-ETs are linked by the function IF.
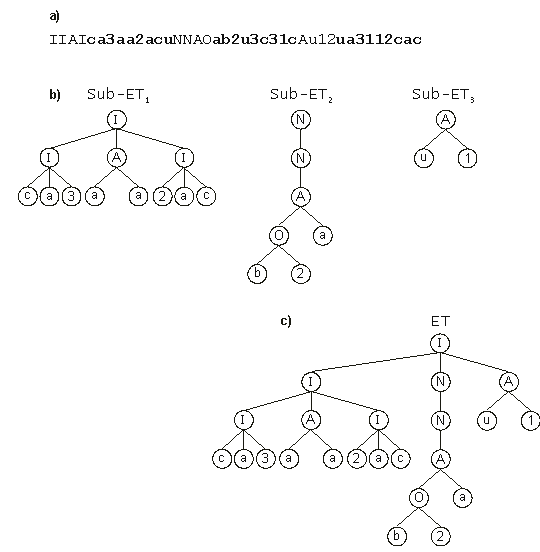
Figure 4. Expression of multigenic chromosomes as ETs. (a) A three-genic chromosome with the tails shown in bold (“N” is a function of one argument and represents NOT; “A” and “O” are functions of two arguments and represent respectively AND and OR; “I” is a function of three arguments and represents IF; the remaining symbols are terminals).
(b) The sub-ETs codified by each gene. (c) The result of posttranslational linking with IF.
Note again that the multi-subunit ET could be linearized as the following K-expression:
|
01234567890123456789012 |
|
|
IINAIAINu1ca3aa2acAOab2 |
(3.11) |
Figure 5 shows another example of sub-ET interaction, where the sub-ETs are of the simplest kind (one-element sub-ETs).
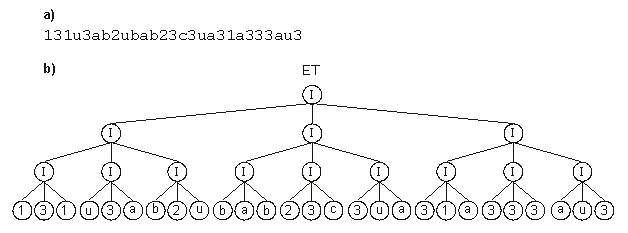
Figure 5. Expression of multigenic chromosomes as ETs. (a) A 27-genic chromosome composed of one-element genes.
(b) The result of posttranslational linking with IF.
In this case, the sub-ETs are linked 3 by 3 with the IF function, then these clusters are, in their turn, linked also 3 by 3 with another IF function, and the three last clusters are also linked by IF, forming a large multi-subunit ET. This kind of chromosomal architecture was used to evolve solutions for the 11-multiplexer problem of
section 6.5.2 and also to evolve cellular automata rules for the density-classification problem. The individual of
Figure 5 could be converted into the following K-expression:
|
0123456789012345678901234567890123456789 |
|
|
IIIIIIIIIIIII131u3ab2ubab23c3ua31a333au3 |
(3.12) |
And finally, the full expression of certain chromosomes requires the sequential execution of small plans, where the first sub-ET does a little work, the second continues from that, and so on. The final plan results from the orderly action of all subplans (see the block stacking problem in
section 6.3).
The type of linking function, as well as the number of genes and the length of each gene, are a priori chosen for each problem. So, we can always start by using a single-gene chromosome, gradually increasing the length of the head; if it becomes very large, we can increase the number of genes and of course choose a function to link them. We can start with addition or OR, but in other cases another linking function might be more appropriate. The idea, of course, is to find a good solution, and GEP provides the means of finding one.
|


